Summary¶
The Modular Autonomous Discovery for Science (MADSci) toolkit enables laboratory automation, high-throughput experimentation, and self-driving labs. MADSci provides a microservices architecture integrating laboratory instruments, robots, and devices with coordination services for workflows, data management, and resource tracking. Users define and execute experiments using Python applications and YAML-based workflow definitions. This modular design separates concerns between device integrators, lab operators, and domain scientists. MADSci supports research in biology, chemistry, materials science, and quantum science.
Statement of Need¶
The lab automation ecosystem is fragmented, with many proprietary, costly, or narrowly domain-specific tools. MADSci provides an open-source, extensible, domain-agnostic toolkit integrating equipment, sensors, and robots as “nodes” with managers for workflows, resources, experiments, logging, and data collection. Built on a microservices architecture with RESTful APIs and Python clients, MADSci leverages standard databases (PostgreSQL, MongoDB, Redis, MiniIO) and open-source libraries (FastAPI, Pydantic, SQLModel).
Commercial platforms such as Chemspeed’s Autosuite Seifrid et al., 2024 and Retisoft Genera Retisoft Inc., 2024 provide comprehensive functionality but operate under proprietary licensing that may limit academic accessibility. Open-source alternatives including AlabOS Fei et al., 2024, ChemOS Sim et al., 2024Roch et al., 2020, and Bluesky Allan et al., 2019 demonstrate strong capabilities within specific domains but exhibit varying cross-disciplinary transferability. Specialized systems such as Polybot Wang et al., 2025Vriza et al., 2023 showcase advanced features tailored to particular applications. The Workcell Execution Interface Vescovi et al., 2023, MADSci’s predecessor, emphasizes instrument modularity but lacks advanced features and microservices principles at the management layer. MADSci provides domain-agnostic laboratory automation while maintaining compatibility with diverse instrumentation and institutional requirements.
Software Architecture and Features¶
MADSci’s microservice architecture enables separation of concerns among system components.
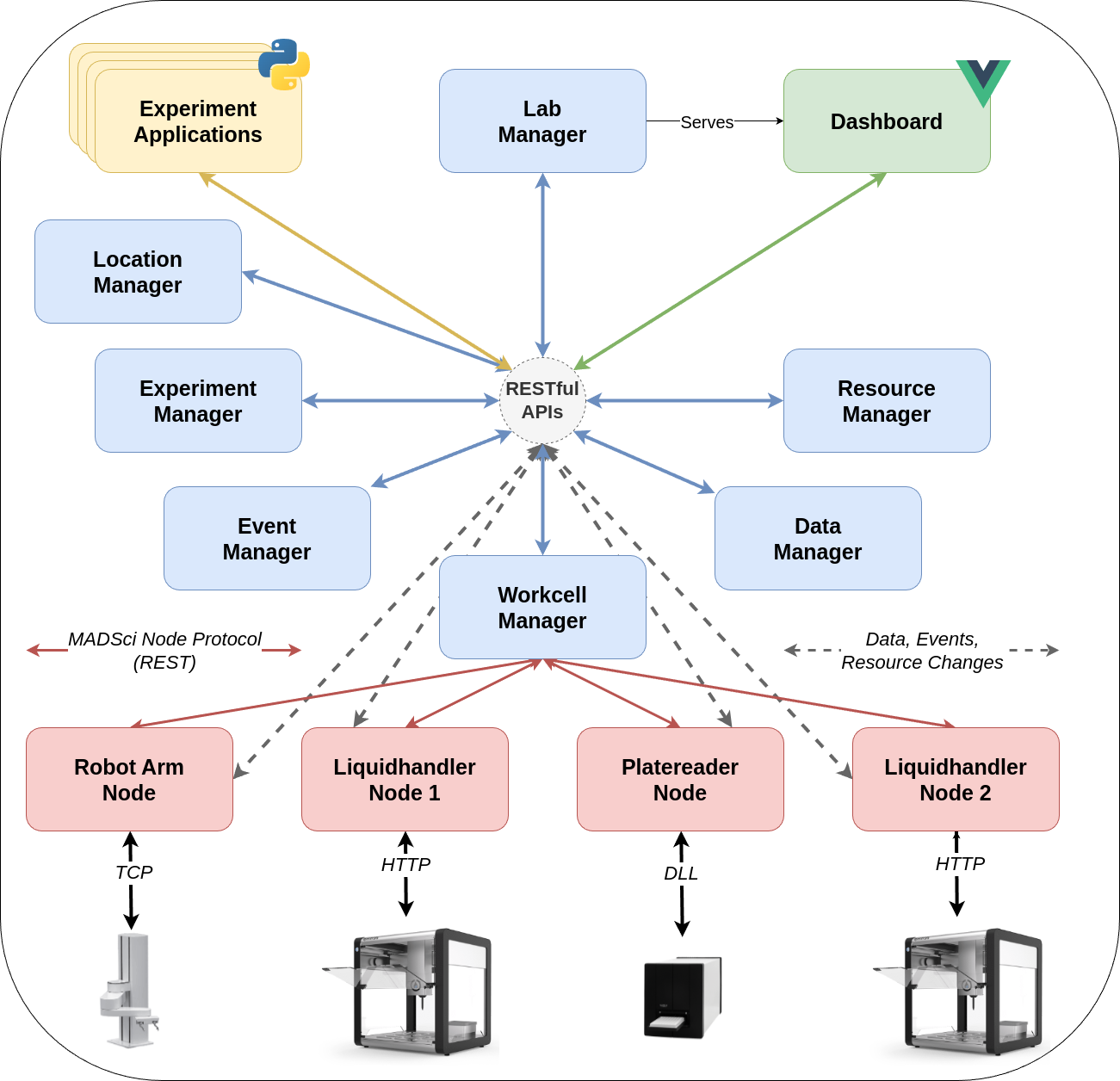
Nodes¶
A node provides a standardized interface to a laboratory instrument, device, robot, or sensor \autoref{fig:madsci_architecture}.
Any software implementing standard API endpoints can function as a node.
We provide a RestNode Python class for device integration.
Key endpoints include /action for device functionality, /status for node state, /state for device information, /admin for administrative commands, and /info for metadata and capabilities.
Workcell and Workflow Management¶
A workcell represents a collection of nodes, physical locations, and resources coordinated to execute experimental procedures. A workflow is a sequence of steps directing nodes to perform specific actions with specified parameters. A self-driving lab may comprise one or more workcells, each executing workflows semi-independently.
The Workcell Manager coordinates operation through a scheduler determining which workflow steps to execute based on current node, location, and resource states. This modular scheduler can be customized by lab operators; we include an opportunistic first-in-first-out scheduler as default.
Workflows are defined using YAML syntax or Pydantic-based Python data models as linear sequences of steps. Each step specifies a target node, an action, and required or optional arguments. Beyond JSON-serializable arguments, steps can reference physical locations or files, with the Workcell Manager resolving location references at runtime.
Workflows support parameterization: users specify node, action, or arguments at submission time, enabling reusable templates. Outputs from earlier steps can inform later parameter values, allowing intra-workflow data flow.
Experiment Management¶
An experiment run represents a single execution of an experimental procedure, tracking associated workflows, resources, data, and metadata. The Experiment Manager enables users to define experiments, initiate runs, and link MADSci objects (workflows, resources, datapoints) with those runs. It supports experiment designs specifying properties and conditions under which experiments are conducted.
The madsci.experiment_application provides classes for defining experimental applications in Python, with client libraries and helper methods for common tasks.
There are helper classes available for writing and running experiment applications as scripts, jupyter notebooks, terminal user interfaces, or even as MADSci nodes themselves, enabling experiment-specific actions within workflows, which can be monitored and managed from the Lab Dashboard.
This design makes laboratory capabilities accessible to domain scientists while remaining flexible for integration with other Python tools.
Data Management¶
The Data Manager supports creation, storage, and querying of data generated during autonomous experimentation.
It stores JSON-serializable data in MongoDB and file-based data on filesystems or S3-compatible object storage.
For large datasets, the DataClient optionally supports direct upload to object storage.
Resource Management¶
Many laboratories, particularly in chemistry and biology, require tracking physical resources such as consumables, labware, and samples. The Resource Manager provides optional capabilities to define, validate, track, and manage these assets across their lifecycle. Built on PostgreSQL, it supports diverse asset types and hierarchical organization with customizable properties and standardized operations. It maintains automated histories and locking mechanisms for provenance and reliability.
Location Management¶
The Location Manager provides optional tracking of physical laboratory locations and their associations with resources. Locations represent positions such as instrument slots, storage areas, or transfer stations within the laboratory environment.
This manager integrates with the Resource Manager to enable attachment of resources to specific locations, facilitating automated tracking of sample positions and consumable storage. Workflows can reference locations symbolically, with the Workcell Manager resolving these references at runtime based on current attachments and states. This abstraction separates physical laboratory layout from workflow logic, improving workflow portability across different laboratory configurations.
Event Management and Logging¶
The Event Manager enables nodes and managers to log JSON events to a central MongoDB-backed system supporting advanced queries.
The EventClient also optionally supports OpenTelemetry-based tracing, metrics, and logging, as well as console- and file-based logging via the structlog Python package, enabling multiple different logging and observability modalities.
Lab Management¶
The Lab Manager provides a primary entrypoint for users and applications. A web-based Dashboard provides overview and detailed information on lab status, performance, and history. The Lab Manager API surfaces context about available managers and lab configuration.
References¶
Acknowledgments¶
The authors acknowledge financial support from Laboratory Directed Research and Development (LDRD) funding from Argonne National Laboratory, provided by the Director, Office of Science, of the U.S. Department of Energy under Contract No. DE-AC02-06CH11357.
- Seifrid, M., Strieth-Kalthoff, F., Hao, H., & others. (2024). Chemspyd: an open-source python interface for Chemspeed robotic chemistry and materials platforms. Digital Discovery. 10.1039/D4DD00046C
- Retisoft Inc. (2024). Genera Laboratory Automation Software. https://retisoft.com/
- Fei, Y., Rendy, B., Kumar, R., & others. (2024). AlabOS: a Python-based reconfigurable workflow management framework for autonomous laboratories. Digital Discovery, 3(11). 10.1039/D4DD00129J
- Sim, M., Pablo-García, S., Schrier, J., & Aspuru-Guzik, A. (2024). ChemOS 2.0: An orchestration architecture for chemical self-driving laboratories. Matter, 7(5), 1871–1895. 10.1016/j.matt.2024.04.021
- Roch, L. M., Häse, F., Kreisbeck, C., & others. (2020). ChemOS: An orchestration software to democratize autonomous discovery. PLOS ONE, 15(4), e0229862. 10.1371/journal.pone.0229862
- Allan, D., Caswell, T., Campbell, S., & Rakitin, M. (2019). Bluesky’s Ahead: A Multi-Facility Collaboration for an a la Carte Software Project for Data Acquisition and Management. Synchrotron Radiation News, 32(3), 19–22. 10.1080/08940886.2019.1608121
- Wang, C., Kim, Y., Vriza, A., & others. (2025). Autonomous platform for solution processing of electronic polymers. Nature Communications, 16, 1498. 10.1038/s41467-024-55655-3
- Vriza, A., Chan, H., & Xu, J. (2023). Self-Driving Laboratory for Polymer Electronics. Chemistry of Materials, 35(8), 3046–3056. 10.1021/acs.chemmater.2c03593
- Vescovi, R., Ginsburg, T., Hippe, K., & others. (2023). Towards a modular architecture for science factories. Digital Discovery. 10.1039/D3DD00142C